Scientific Publications
filter by journal
our timeline of articles
PEER REVIEWED JOURNALS

Reversible Light/Dark Modulation of Spinach Leaf Nitrate Reductase Activity involves Protein Phosphorylation
Spinach (Spinacia oleracea L.) leaf nitrate reductase (NADH:NR;NADH:nitrate oxidoreductase, EC 1.6.6.1) activity was found to rapidly change during light/dark transitions. The most rapid and dramatic changes were found in a form of NR which was sensitive to inhibition by millimolar concentrations of magnesium. This form of NR predominated in leaves in the dark, but was almost completely absent from leaves incubated in the light for only 30 min. When the leaves were returned to darkness, the NR rapidly became sensitive to Mg2+ inhibition. Modulation of the overall reaction involving NADH as electron donor was also found when reduced methyl viologen was the donor (MV:NR), indicating that electron transfer had been blocked, at least in part, at or near the terminal molybdenum cofactor site. Changes in activity appear to be the result of a covalent modification that affects sensitivity of NR to inhibition by magnesium, and our results suggest that protein phosphorylation may be involved. NR was phosphorylated in vivo after feeding excised leave [32P]Pi. The NR subunit was labeled exclusively on seryl residues in both light and dark. Tryptic peptide mapping indicated three major 32P-labeled phosphopeptide (Pp) fragments. Labeling of two of the P-peptides (designated Pp1 and 3) was generally correlated with NR activity assayed in the presence of Mg2+. In vivo, partial dephosphorylation of these sites (and activation of NR assayed with Mg2+) occurred in response to light or feeding mannose in darkness. The light effect was blocked completely by feeding okadaic acid via the transpiration stream, indicating the involvement of type 1 and/or type 2A protein phosphatases in vivo. While more detailed analysis is required to establish a causal link between the phosphorylation status of NR and sensitivity to Mg2+ inhibition, the current results are highly suggestive of one. Thus, in addition to the molecular genetic mechanisms regulating this key enzyme of nitrate assimilation, NR activity may be controlled in leaves by phosphorylation/ dephosphorylation of the enzyme protein resulting from metabolic changes taking place during light/dark transitions.View Full Article
Field Determination of Nitrate using Nitrate Reductase: Proceeding of the Symposium on Field Analytical Methods for Hazardous Wastes and Toxic Chemicals: A&WMA Conference
Nitrate is routinely measured in a variety of substrates - water, tissues, soils, and foods - both in the field and in laboratory settings. The most commonly used nitrate test methods involve the reduction of nitrate to nitrite via a copper-cadmium reagent, followed by reaction of the nitrite with the Griess dye reagents. The resulting color is translated into a nitrate concentration by comparison with a calibrated color chart or comparator, or by reading the absorbance in a spectrophotometer. This basic method is reliable and sufficiently sensitive for many applications. However, the cadmium reagent is quite toxic. The trend today is for continued increase in concern for worker health and safety; in addition, there are increasing costs and logistical problems associated with regulatory constraints on transport and disposal of hazardous materials. Some suppliers have substituted a zinc-based nitrate detection method as an improvement on the basic Griess method that demonstrates equal or superior sensitivity, superior selectivity, and is more environmentally benign. Comparisons between the enzyme-based method and some standard field test kits being used today are made. View Full Article
Construction and Characterization of Nitrate Reductase-Based Amperometric Electrode and Nitrate Assay of Fertilizers and Drinking Water
The construction and characterization of a nitrate reductase-based amperometric electrode for determination of nitrate ion is described. The electrode consisted of nitrate reductase held by dialysis membrane onto a Nafion-coated glassy carbon electrode. Methyl viologen was allowed to absorb into the Nafion layer, which acted as a reservoir for the electron mediator. The utility of the electrode to assay fertilizer and water sample for nitrate was demonstrated. The assays conducted with this electrode compared well with colorimetric and potentiometric assays of the same samples. View Full Article
Nitrate Reductase Structure, Function and Regulation: Bridging the Gap between Biochemistry and Physiology
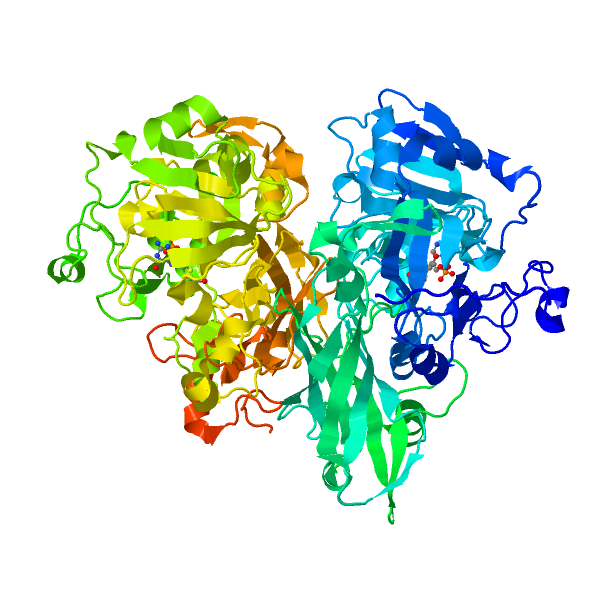
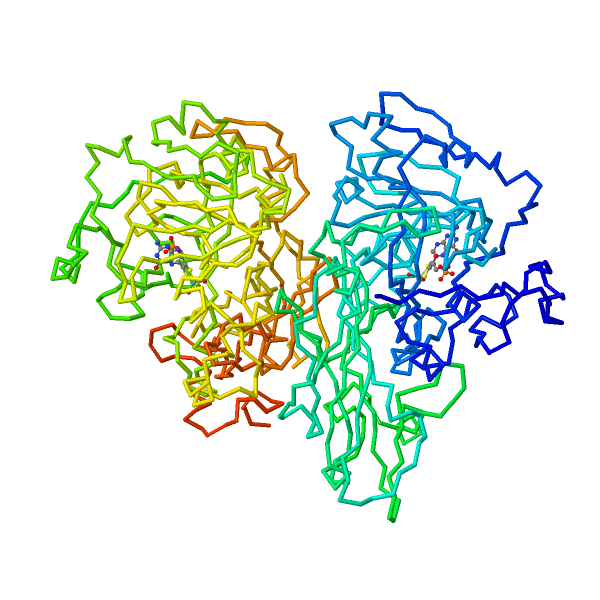
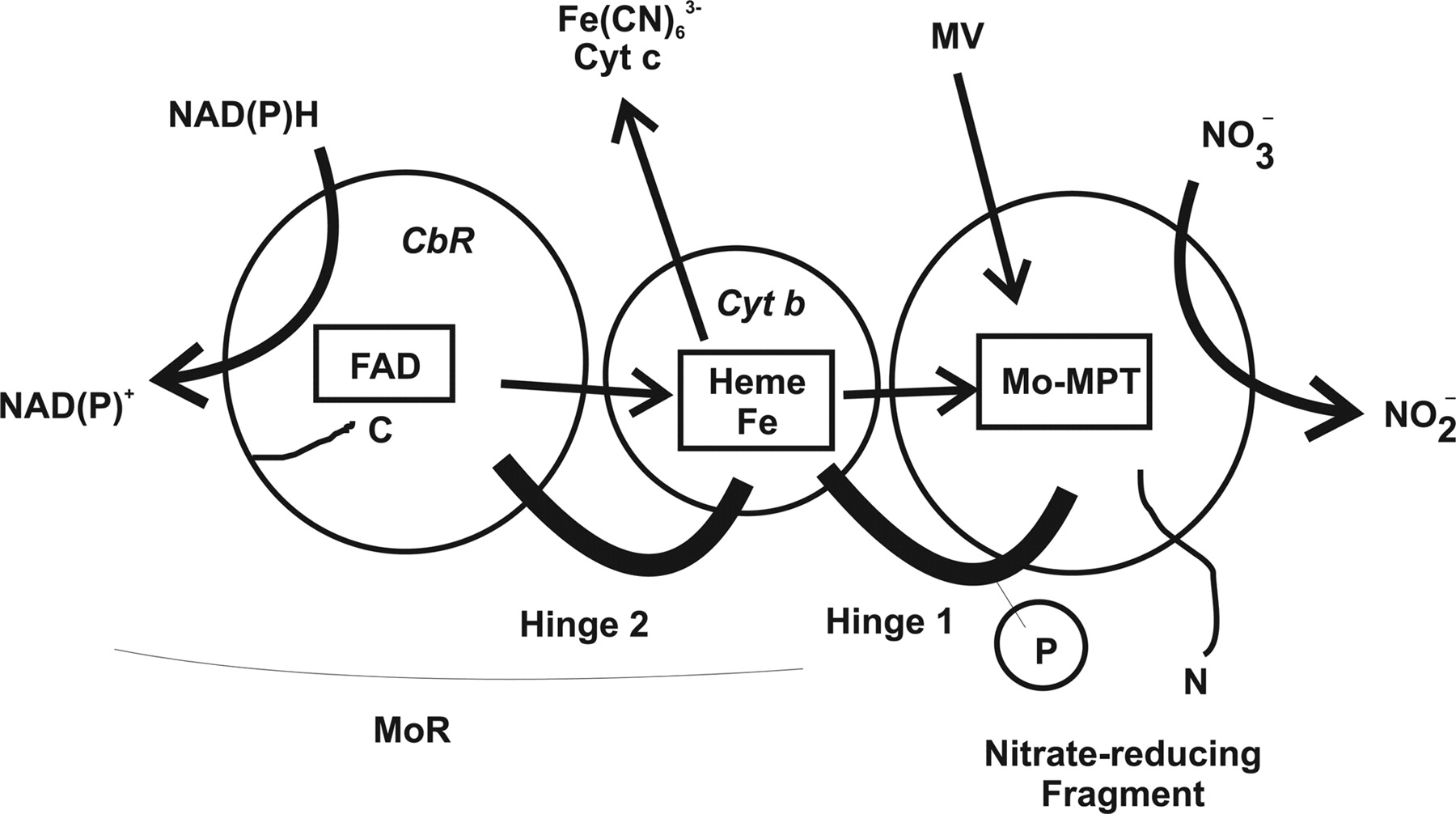
Nitrate reductase (NR; EC 1.6.6.1-3) catalyzes NAD(P)H reduction of nitrate to nitrite. NR serves plants, algae, and fungi as a central point for integration of metabolism by governing flux of reduced nitrogen by several regulatory mechanisms. The NR monomer is composed of a ∼100-kD polypeptide and one each of FAD, heme-iron, and molybdenum-molybdopterin (Mo-MPT). NR has eight sequence segments: (a) N-terminal “acidic” region; (b) Mo-MPT domain with nitrate-reducing active site; (c) interface domain; (d) Hinge 1 containing serine phosphorylated in reversible activity regulation with inhibition by 14-3-3 binding protein; (e) cytochrome b domain; (f) Hinge 2; (g) FAD domain; and (h) NAD(P)H domain. The cytochrome b reductase fragment contains the active site where NAD(P)H transfers electrons to FAD. A complete three-dimensional dimeric NR structure model was built from structures of sulfite oxidase and cytochrome b reductase. Key active site residues have been investigated. NR structure, function, and regulation are now becoming understood. View Full Article
Development, characterization, and operational details of an enzymatic, air-segmented continuous-flow analytical method for colorimetric determination of nitrate + nitrite in natural-water samples is described. This method is similar to U.S. Environmental Protection Agency method 353.2 and U.S. Geological Survey method I-2545-90 except that nitrate is reduced to nitrite by soluble nitrate reductase (NaR, EC 1.6.6.1) purified from corn leaves rather than a packed-bed cadmium reactor. A three-channel, air-segmented continuous-flow analyzerconfigured for simultaneous determination of nitrite (0.020−1.000 mg-N/L) and nitrate + nitrite (0.05−5.00 mg-N/L) by the nitrate reductase and cadmium reduction methodswas used to characterize analytical performance of the enzymatic reduction method. At a sampling rate of 90 h-1, sample interaction was less than 1% for all three methods. Method detection limits were 0.001 mg of NO2- -N/L for nitrite, 0.003 mg of NO3-+ NO2- -N/L for nitrate + nitrite by the cadmium-reduction method, and 0.006 mg of NO3-+ NO2- -N/L for nitrate + nitrite by the enzymatic-reduction method. Reduction of nitrate to nitrite by both methods was greater than 95% complete over the entire calibration range. The difference between the means of nitrate + nitrite concentrations in 124 natural-water samples determined simultaneously by the two methods was not significantly different from zero at the p = 0.05 level.View Full Article
Nitrate analysis in water is one of the most frequently applied methods in environmental chemistry. Current methods for nitrate are generally based on toxic substances. Here, we show that a viable alternative method is to use the enzyme nitrate reductase. The key to applying this Green Chemistry solution for nitrate analysis is plentiful, inexpensive, analytical grade enzyme. We demonstrate that recombinant Arabidopsis nitrate reductase, expressed in the methylotrophic yeast Pichia pastoris, is a highly effective catalyst for nitrate analysis at 37°C. Recombinant production of enzyme ensures consistent quality and provides means to meet the needs of environmental chemistry.View Full Article
Genetic Modification of Pichia Pastoris for Production of Propylene Glycol from Glycerol
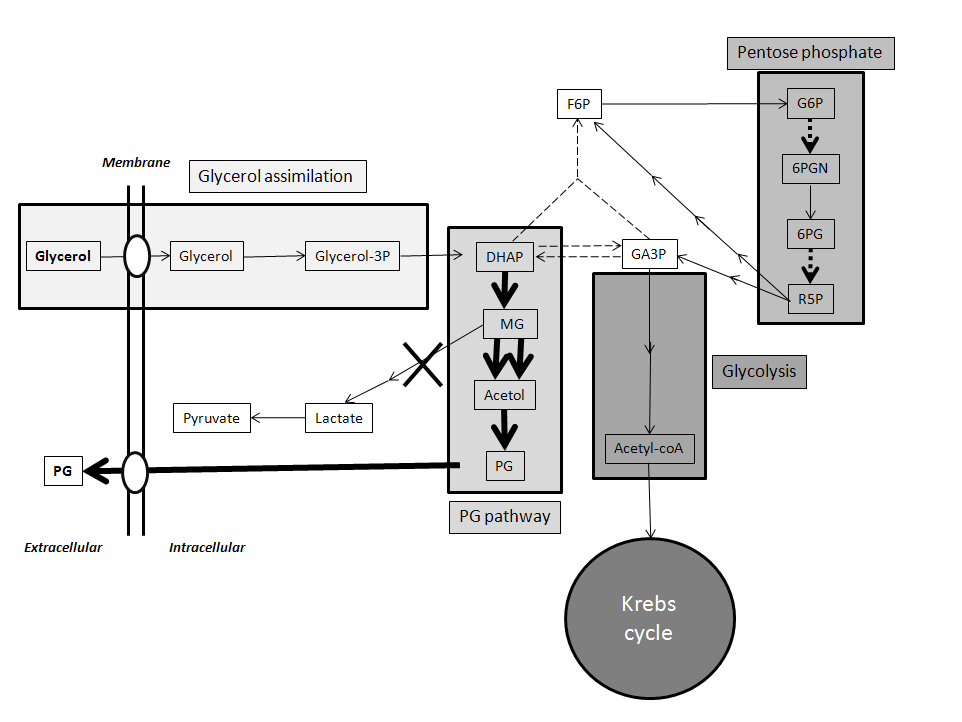
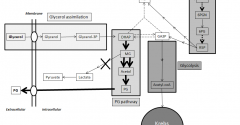
Biodiesel is emerging as a major renewable energy resource. Glycerol is byproduct of biodiesel production, which cannot be directly used as fuel. Using Pichia pastoris as aerobic yeast cell factory, we engineered pathway to convert glycerol to propylene glycol (PG; 1,2-propanediol, 1,2-dihydroxypropane), a fuel additive and commodity compound. PG biosynthetic pathway was designed with unique combination of 3 enzymes: Escherichia coli methylglyoxal synthase and alcohol dehydrogenase; and Pichia ofunaensis glycerol dehydrogenase. Engineered genes were successfully integrated in yeast genome and actively expressed. Analysis, by HPLC of shake flask and fermenter broths of glycerol-grown cells, demonstrated engineered yeast produced PG. In fermenter, highest PG concentration was 0.11 g/L. When engineered P. pastoris was grown on biodiesel crude glycerol as sole carbon source, PG was detected. Basically, this is a proof of concept to demonstrate that Pichia was engineered to produce PG from glycerol.View Full Article
Enzyme-Catalyzed O2 Removal System for Electrochemical Analysis Under Ambient Air: Application in an Ampherometric Nitrate Biosensor
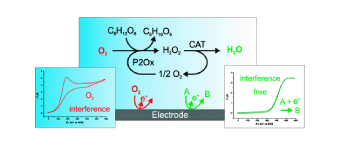
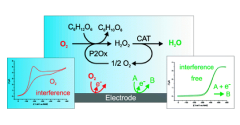
Electroanalytical procedures are often subjected to oxygen interferences. However, achieving anaerobic conditions in field analytical chemistry is difficult. In this work, novel enzymatic systems were designed to maintain oxygen-free solutions in open, small volume electrochemical cells and implemented under field conditions. The oxygen removal system consists of an oxidase enzyme, an oxidase-specific substrate, and catalase for dismutation of hydrogen peroxide generated in the enzyme catalyzed oxygen removal reaction. Using cyclic voltammetry, three oxidase enzyme/substrate combinations with catalase were analyzed: glucose oxidase with glucose, galactose oxidase with galactose, and pyranose 2-oxidase with glucose. Each system completely removed oxygen for 1 h or more in unstirred open vessels. Reagents, catalysts, reaction intermediates, and products involved in the oxygen reduction reaction were not detected electrochemically. To evaluate the oxygen removal systems in a field sensing device, a model nitrate biosensor based on recombinant eukaryotic nitrate reductase was implemented in commercial screen-printed electrochemical cells with 200 μL volumes. The products of the aldohexose oxidation catalyzed by glucose oxidase and galactose oxidase deactivate nitrate reductase and must be quenched for biosensor applications. For general application, the optimum catalyst is pyranose 2-oxidase since the oxidation product does not interfere with the biorecognition element. View Full Article
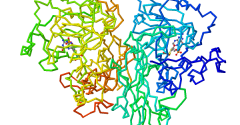

Affinity binding via Zinc(II) for controlled orientation and electrochemistry of Histidine-tagged nitrate reductase in self-assembled monolayers
Monolayers of CuII-complexes on electrode surfaces are frequently applied for the immobilization and controlled orientation of His-tagged redox proteins. However, affinity binding is limited to applications that require potentials less negative than the reduction potential of the metal complexes. In order to overcome this limitation, we used Zn2 + cations on nitrilotriacetic acid (NTA) modified carbon electrodes for the coordination of His-tagged nitrate reductase (NaR). The NTA modified electrodes were prepared upon diazotation and electrochemical reduction of an aniline functionalized NTA ligand. After coordination of Zn2 + to the bound NTA ligand, self-assembly of NaR is achieved via coordination of the imidazole groups from the His-tag to the NTA–ZnII complex. The electrochemical investigations of the NaR monolayer on NTA–ZnII films demonstrate the catalytic activity for reduction of nitrate to nitrite in the presence of methyl viologen. The catalytic current density correlates with the one expected for a fully active enzyme monolayer. Moreover, the reduction of Zn2 + is not observed at the potential necessary for the reduction of methyl viologen. Therefore, affinity binding based on Zn2 + may be used for the immobilization and electrochemical applications of His-tagged NaR.
Determination of Phosphate in Soil Extracts in the Field: A Green Chemistry Enzymatic Method
Measurement of ortho-phosphate in soil extracts usually involves sending dried samples of soil to a laboratory for analysis and waiting several weeks for the results. Phosphate determination methods often involve use of strong acids, heavy metals, and organic dyes. To overcome limitations of this approach, we have developed a phosphate determination method which can be carried out in the field to obtain results on the spot.
This new method uses:
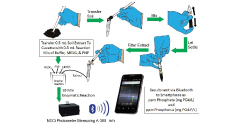
First, the soil sample is extracted with deionized water and filtered. Next, an aliquot of the soil extract (0.5 mL) is transferred to a disposable cuvette, containing 0.5 mL of reaction mixture [200 mM HEPES, pH 7.6, 20 mM MgCl2, with 80 nmol 2-amino-6-mercapto-7-methylpurine ribonucleoside (MESG) and 1 unit of recombinant purine nucleoside phosphorylase (PNP; EC 2.4.2.1)], mixed, and incubated for 10 min at field temperature. Absorbance of the completed reaction is measured at 360 nm in open-source, portable photometer linked by bluetooth to a smartphone. The phosphate and phosphorus content of the soil is determined by comparison of its absorbance at 360 nm to a previously prepared standard phosphate curve, which is stored in the smartphone app. View the full article

Non-toxic total nitrogen determination using a low alkaline persulfate digestion
Non-toxic total nitrogen determination using a low alkaline persulfateRead more
Read moreMeasuring total nitrogen, nitrate, and nitrite is critical for compliance with water safety standards. Previous methods for measuring total nitrogen were hazardous, time consuming, and expensive. Here we report a method for measuring total nitrogen in water and soil using alkaline persulfate digestion combined with a Nitrate Reductase assay. In this method the alkaline persulfate reaction oxidizes all nitrogen present in the sample to nitrate, Nitrate Reductase then is used to catalyze the reduction of nitrate to nitrite in the presence of NADH. The nitrite is then treated with Griess reagents to produce a pink color. The absorbance of this color is measured at 540 nm using a spectrophotometer and when compared to a standard curve of nitrate, treated with both the reduction and colorizing steps, can be used to determine the total nitrogen content of measured samples. This method customizes the measurement of total nitrogen by combining alkaline persulfate digestion with a Nitrate Reductase assay using enzyme based green chemistry. View Full Article